Run a Real Simulation
Summary
This module aims to do a parameter study on a well-known benchmark problem for viscous incompressible fluid flow.
Prerequisites
Estimated Time
60 minutes
You Will learn
- How to run the simulation OpenFOAM, using Merlin.
- How to use machine learning on OpenFOAM results, using Merlin.
Introduction
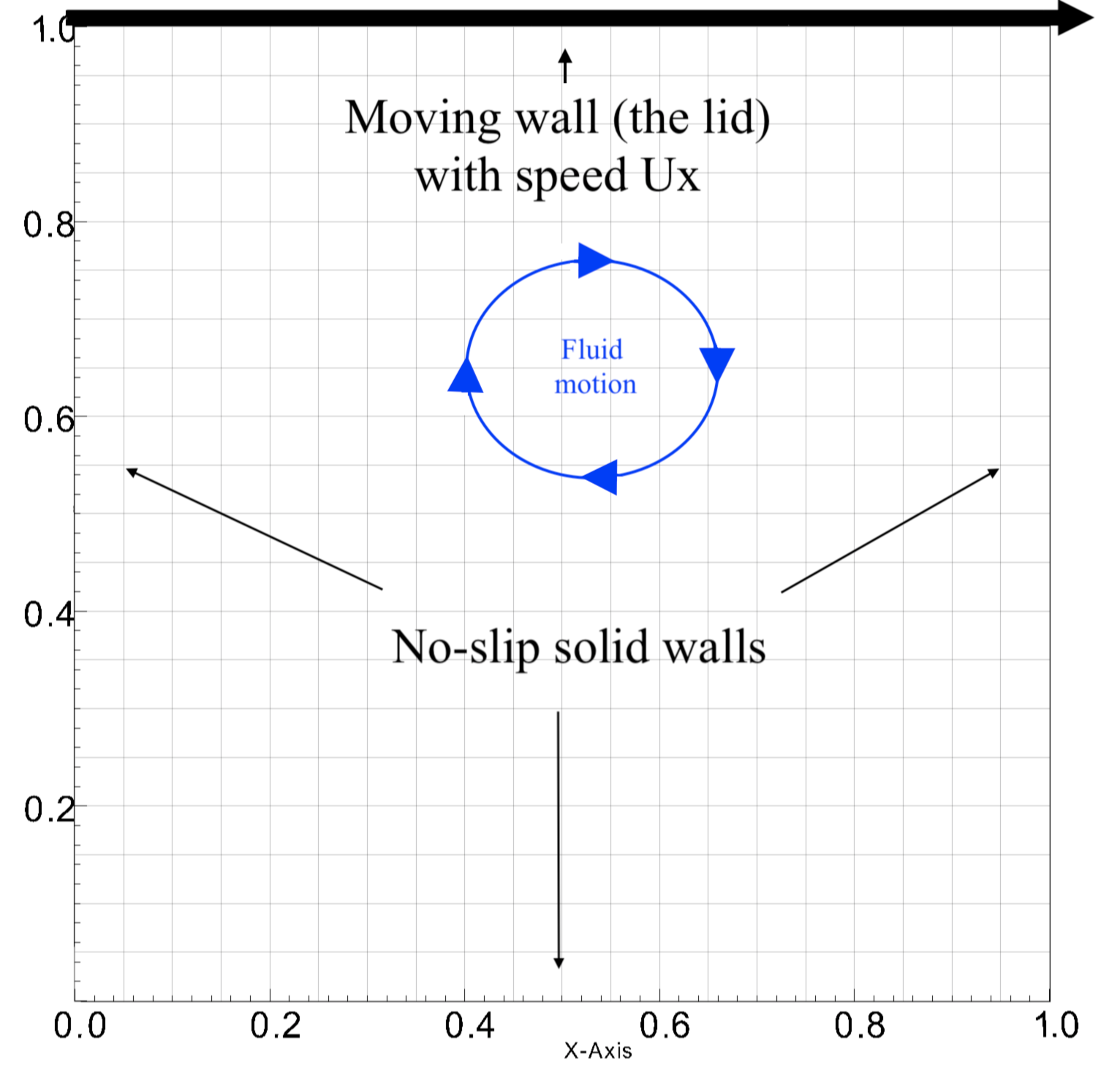
We aim to do a parameter study on the lid-driven cavity problem.
 |
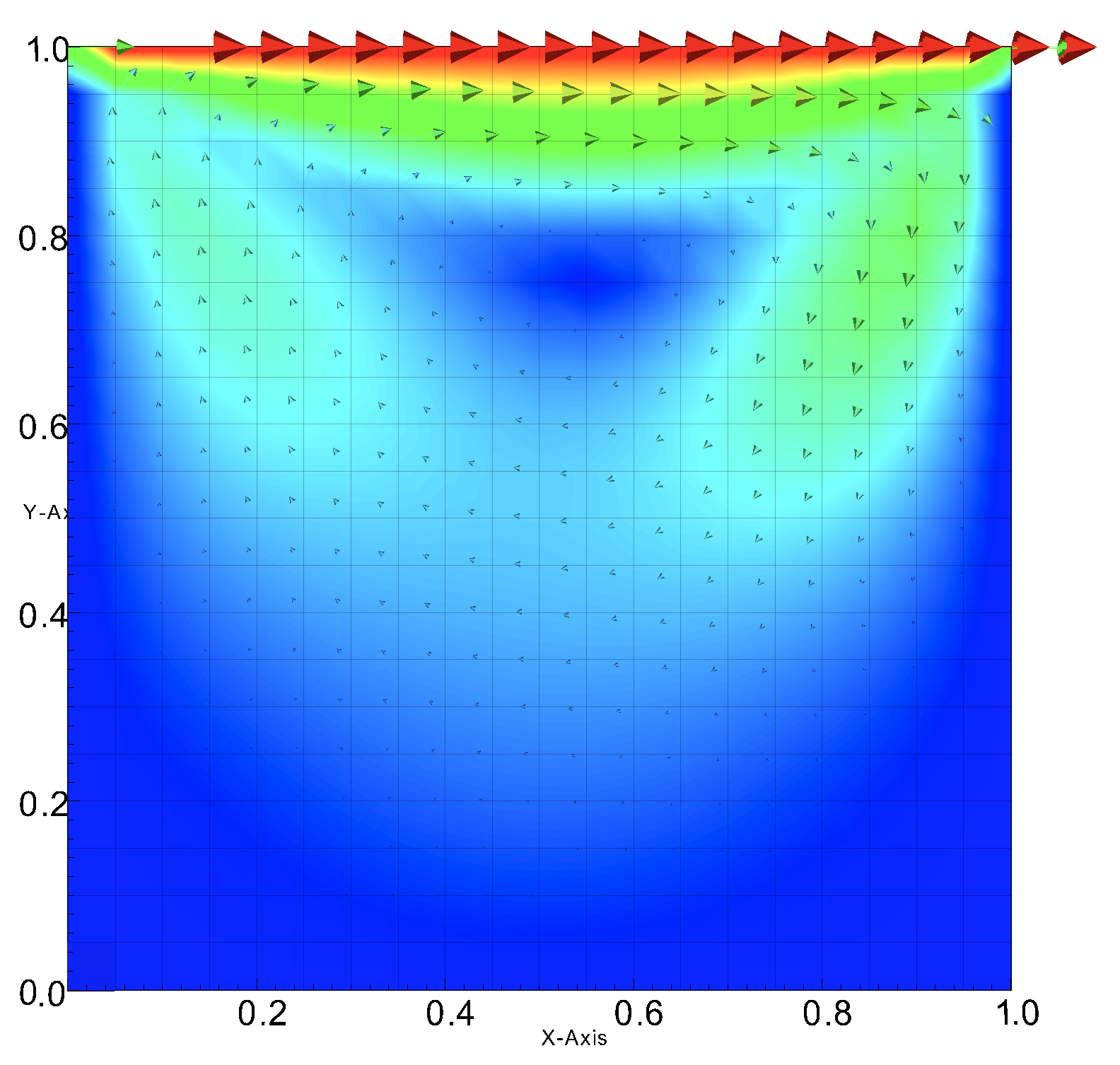 |
In this problem, we have a viscous fluid within a square cavity that has three non-slip walls and one moving wall (moving lid). We are interested in learning how varying the viscosity and lid speed affects the average enstrophy and kinetic energy of the fluid after it reaches steady state. We will be using the velocity squared as a proxy for kinetic energy.
This module will be going over:
- Setting up our inputs using the
merlinblock - Running multiple simulations in parallel
- Combining the outputs of these simulations into a an array
- Predictive modeling and visualization
Before Moving On
Check that the virtual environment with Merlin installed is activated and that your configuration shows no errors:
This is covered more in depth in the Pointing Merlin to the Server section of the Installation module and at the Configuration page.
There are a few ways to do this example, including with singularity and with docker. To go through the version with singularity, get the necessary files for this module by running:
For the version with docker you should run:
Note
From here on, this tutorial will focus solely on the singularity version of running OpenFOAM. The docker version of this tutorial is almost identical but will require having docker installed. If you're using the docker version of this tutorial you can still follow along but check the openfoam_docker_template.yaml file in each step to see what differs.
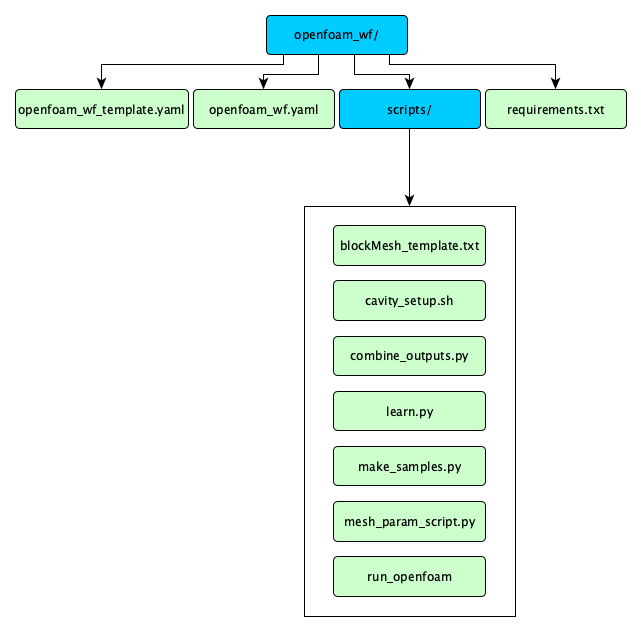
In the openfoam_wf_singularity directory you should see the following:
openfoam_wf_singularity.yaml-- this spec file is partially blank. You will fill in the gaps as you follow this module's steps.openfoam_wf_singularity_template.yaml-- this is a complete spec file. You can always reference it as an example.scripts-- This directory contains all the necessary scripts for this module.- We'll be exploring these scripts as we go with the tutorial.
requirements.txt-- this is a text file listing this workflow's python dependencies.
Specification File
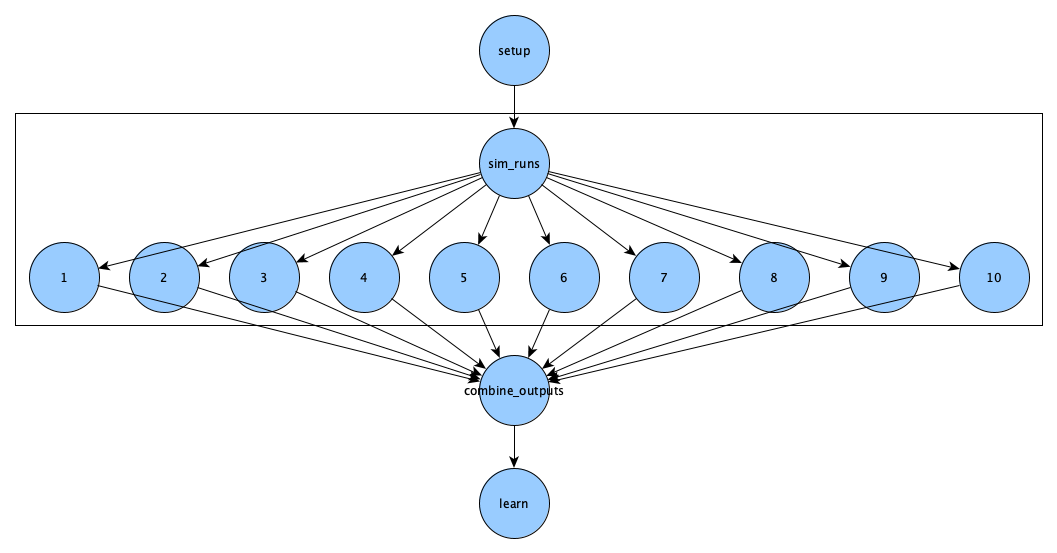
We are going to build a spec file that produces this DAG:
To start, open openfoam_wf_singularity.yaml using your favorite text editor.
It should look something like this:
Initial Contents of the Spec
description:
name: openfoam_wf_singularity
description: |
A parameter study that includes initializing, running,
post-processing, collecting, learning and visualizing OpenFOAM runs
using singularity.
env:
variables:
OUTPUT_PATH:
SCRIPTS:
SIF:
N_SAMPLES:
merlin:
samples:
generate:
cmd: |
file:
column_labels:
resources:
workers:
nonsimworkers:
args: -l INFO --concurrency <INPUT CONCURRENCY HERE>
steps:
simworkers:
args: -l INFO --concurrency <INPUT CONCURRENCY HERE> --prefetch-multiplier 1 -Ofair
steps:
study:
- name: setup
description: |
Installs necessary python packages and imports the cavity directory
from the singularity container
run:
cmd: |
- name: sim_runs
description: |
Edits the Lidspeed and viscosity then runs OpenFOAM simulation
using the icoFoam solver
run:
cmd: |
depends:
task_queue: simqueue
- name: combine_outputs
description: Combines the outputs of the previous step
run:
cmd: |
depends:
- name: learn
description: Learns the output of the openfoam simulations using input parameters
run:
cmd: |
depends:
Variables
First we specify some variables to make our life easier. Locate the env block in our yaml spec:
The OUTPUT_PATH variable is set to tell Merlin where you want your output directory to be written. The default is the current working directory.
We'll fill out the next two variables as we go.
Samples and Scripts
One Merlin best practice is to copy any scripts your workflow may use from your SPECROOT directory into the MERLIN_INFO directory. This is done to preserve the original scripts in case they are modified during the time Merlin is running. We will do that first. We'll put this in the Merlin sample generation section, since it runs before anything else.
Edit the samples section of the merlin block to look like the following:
merlin:
samples:
generate:
cmd: |
cp -r $(SPECROOT)/scripts $(MERLIN_INFO)/
# Generates the samples
python $(MERLIN_INFO)/scripts/make_samples.py -n 10 -outfile=$(MERLIN_INFO)/samples
file: $(MERLIN_INFO)/samples.npy
column_labels: [LID_SPEED, VISCOSITY]
We will be using the scripts directory a lot so we'll set the variable SCRIPTS to $(MERLIN_INFO)/scripts for convenience. We define SIF, our Singularity image of OpenFoam, to be $(MERLIN_INFO)/openfoam6.sif. We would also like to have a more central control over the number of samples generated so we'll create an N_SAMPLES variable:
env:
variables:
OUTPUT_PATH: ./openfoam_wf_output
SCRIPTS: $(MERLIN_INFO)/scripts
SIF: $(MERLIN_INFO)/openfoam6.sif
N_SAMPLES: 100
and update the samples section of the merlin block to be:
merlin:
samples:
generate:
cmd: |
cp -r $(SPECROOT)/scripts $(MERLIN_INFO)/
# Generates the samples
python $(SCRIPTS)/make_samples.py -n $(N_SAMPLES) -outfile=$(MERLIN_INFO)/samples
file: $(MERLIN_INFO)/samples.npy
column_labels: [LID_SPEED, VISCOSITY]
Just like in the Using Samples step of the hello world module, we generate samples using the merlin block. We are only concerned with how the variation of two initial conditions, lidspeed and viscosity, affects outputs of the system. These are the column_labels. The make_samples.py script is designed to make log uniform random samples. Now, we can move on to the steps of our study block.
Setting Up
Our first step in our study block is concerned with making sure we have all the required python packages for this workflow. The specific packages are found in the requirements.txt file.
We will also need to copy the lid driven cavity deck from the OpenFOAM singularity container and adjust the write controls. This last part is scripted already for convenience.
Locate the setup step in the study block and edit it to look like the following:
study:
- name: setup
description: |
Installs necessary python packages and imports the cavity directory
from the singularity container
run:
cmd: |
pip install -r $(SPECROOT)/requirements.txt
# Set up the cavity directory in the MERLIN_INFO directory
source $(SCRIPTS)/cavity_setup.sh $(MERLIN_INFO)
This step does not need to be parallelized so we will assign it to lower concurrency (a setting that controls how many workers can be running at the same time on a single node).
Locate the resources section in the merlin block, then edit the concurrency and add the setup step:
The resources section of the merlin block is where you can control the behavior of your workers. Here we're defining one worker named nonsimworkers and providing it with some arguments. The -l INFO option sets its log level to be INFO (the standard log level) and the --concurrency 1 option means that only one of these workers can spin up per node.
Warning
If the --concurrency option is omitted, the Celery library will default to using however many cores are on the node (see Celery's docs for more information).
In addition to providing arguments to workers, we're also telling it to manage the tasks that will be produced by the setup step. What this actually means is that this worker will watch the task_queue associated with the setup step. Since we didn't provide a task_queue value for this step, the task_queue value will default to be merlin (which you'll be able to see in the openfoam_wf.expanded.yaml spec in the merlin_info/ directory after we run this).
Running the Simulation
Moving on to the sim_runs step, we want to:
- Copy the cavity deck from the
MERLIN_INFOdirectory into each of the current step's subdirectories - Edit the default input values (lidspeed and viscosity) in these cavity decks using the
sedcommand - Run the simulation using the
run_openfoamexecutable through the OpenFOAM singularity container - Post-process the results (also using the
run_openfoamexecutable)
This part should look like:
- name: sim_runs
description: |
Edits the Lidspeed and viscosity then runs OpenFOAM simulation
using the icoFoam solver
run:
cmd: |
cp -r $(MERLIN_INFO)/cavity cavity/
cd cavity
## Edits default values for viscosity and lidspeed with
# values specified by samples section of the merlin block
sed -i "18s/.*/nu [0 2 -1 0 0 0 0] $(VISCOSITY);/" constant/transportProperties
sed -i "26s/.*/ value uniform ($(LID_SPEED) 0 0);/" 0/U
cd ..
cp $(SCRIPTS)/run_openfoam .
# The local sample directory is bound (--bind) to the /merlin_sample dir in
# the container instance
# The sample/cavity is bound (--bind) to the /cavity dir in the container instance
REPATH=$(WORKSPACE)/$(MERLIN_SAMPLE_PATH)
CBIND=${REPATH}:/merlin_sample,${REPATH}/cavity:/cavity
# Creating a unique OpenFOAM singularity execution for each sample to run the simulation
singularity exec --bind ${CBIND} $(SIF) /merlin_sample/run_openfoam $(LID_SPEED)
depends: [setup]
task_queue: simqueue
This step runs many simulations in parallel so it would run faster if we assign it a worker with a higher concurrency. Navigate back to the resources section in the merlin block
resources:
workers:
nonsimworkers:
args: -l INFO --concurrency 1
steps: [setup]
simworkers:
args: -l INFO --concurrency 10 --prefetch-multiplier 1 -Ofair
steps: [sim_runs]
Since we defined a task_queue value in the sim_runs step, when we tell simworkers to watch the sim_runs step they will really be watching the simqueue queue.
The quantities of interest are the average enstrophy and kinetic energy at each cell. The enstrophy is calculated through an OpenFOAM post processing function of the the flow fields while the kinetic energy is approximated by using the square of the velocity vector at each grid point. The velocity field is normally outputted normally as a result of running the default solver for this particular problem.
The run_openfoam executable calculates the appropriate timestep deltaT so that we have a Courant number of less than 1. It also uses the icoFoam solver on the cavity decks and gives us VTK files that are helpful for visualizing the flow fields using visualization tools such as VisIt or ParaView.
Combining Outputs
Navigate to the next step in our study block combine_outputs. The purpose of this step is to extract the data from each of the simulation runs from the previous step (sim_runs) and combine them for future use.
The combine_outputs.py script in the $(SCRIPTS) directory is provided for convenience and takes two inputs. The first informs it of the base directory of the sim_runs directory and the second specifies the subdirectories for each run. The script then goes into each of the directories and combines the velocity and enstrophy for each timestep of each run in a .npz file.
- name: combine_outputs
description: Combines the outputs of the previous step
run:
cmd: |
python $(SCRIPTS)/combine_outputs.py -data $(sim_runs.workspace) -merlin_paths $(MERLIN_PATHS_ALL)
depends: [sim_runs_*]
The $(MERLIN_PATHS_ALL) variable is a Reserved Variable that denotes a space delimited string of all of the sample paths.
This step depends on all the previous step's simulation runs which is why we have the _*. However, it does not need to be parallelized so we assign it to the nonsimworkers in the workers section of the merlin block.
Machine Learning and Visualization
In the learn step, we want to:
- Post-process the .npz file from the previous step.
- Learn the mapping between our inputs and chosen outputs
- Graph important features
The provided learn.py script does all of the above. It outputs the trained sklearn model and a png of the graphs plotted in the current directory.
- name: learn
description: Learns the output of the openfoam simulations using input parameters
run:
cmd: |
python $(SCRIPTS)/learn.py -workspace $(MERLIN_WORKSPACE)
depends: [combine_outputs]
This step is also dependent on the previous step for the .npz file and will only need one worker therefore we will assign it to nonsimworkers:
Putting It All Together
By the end, your openfoam_wf_singularity.yaml should look like the template version in the same directory:
Complete Spec File
description:
name: openfoam_wf_singularity
description: |
A parameter study that includes initializing, running,
post-processing, collecting, learning and visualizing OpenFOAM runs
using singularity.
env:
variables:
OUTPUT_PATH: ./openfoam_wf_output
SCRIPTS: $(MERLIN_INFO)/scripts
SIF: $(MERLIN_INFO)/openfoam6.sif
N_SAMPLES: 100
merlin:
samples:
generate:
cmd: |
cp -r $(SPECROOT)/scripts $(MERLIN_INFO)/
# Generates the samples
python $(SCRIPTS)/make_samples.py -n $(N_SAMPLES) -outfile=$(MERLIN_INFO)/samples
file: $(MERLIN_INFO)/samples.npy
column_labels: [LID_SPEED, VISCOSITY]
resources:
workers:
nonsimworkers:
args: -l INFO --concurrency 1
steps: [setup, combine_outputs, learn]
simworkers:
args: -l INFO --concurrency 10 --prefetch-multiplier 1 -Ofair
steps: [sim_runs]
study:
- name: setup
description: |
Installs necessary python packages and imports the cavity directory
from the singularity container
run:
cmd: |
pip install -r $(SPECROOT)/requirements.txt
# Set up the cavity directory in the MERLIN_INFO directory
source $(SCRIPTS)/cavity_setup.sh $(MERLIN_INFO)
- name: sim_runs
description: |
Edits the Lidspeed and viscosity then runs OpenFOAM simulation
using the icoFoam solver
run:
cmd: |
cp -r $(MERLIN_INFO)/cavity cavity/
cd cavity
## Edits default values for viscosity and lidspeed with
# values specified by samples section of the merlin block
sed -i "18s/.*/nu [0 2 -1 0 0 0 0] $(VISCOSITY);/" constant/transportProperties
sed -i "26s/.*/ value uniform ($(LID_SPEED) 0 0);/" 0/U
cd ..
cp $(SCRIPTS)/run_openfoam .
# The local sample directory is bound (--bind) to the /merlin_sample dir in
# the container instance
# The sample/cavity is bound (--bind) to the /cavity dir in the container instance
REPATH=$(WORKSPACE)/$(MERLIN_SAMPLE_PATH)
CBIND=${REPATH}:/merlin_sample,${REPATH}/cavity:/cavity
# Creating a unique OpenFOAM singularity execution for each sample to run the simulation
singularity exec --bind ${CBIND} $(SIF) /merlin_sample/run_openfoam $(LID_SPEED)
depends: [setup]
task_queue: simqueue
- name: combine_outputs
description: Combines the outputs of the previous step
run:
cmd: |
python $(SCRIPTS)/combine_outputs.py -data $(sim_runs.workspace) -merlin_paths $(MERLIN_PATHS_ALL)
depends: [sim_runs_*]
- name: learn
description: Learns the output of the openfoam simulations using input parameters
run:
cmd: |
python $(SCRIPTS)/learn.py -workspace $(MERLIN_WORKSPACE)
depends: [combine_outputs]
Run the workflow
Now that you are done with the Specification file, use the following commands from inside the openfoam_wf_singularity directory to run the workflow on our task server.
Note
Running with fewer samples is the one of the best ways to debug
Create the DAG and send tasks to the server with:
Open a new terminal window, then start the workers that will consume the tasks we just queued by using:
But wait! We realize that 10 samples is not enough to train a good model. We would like to restart with 100 samples instead of 10 (should take about 6 minutes):
After sending the workers to start on their queues, we need to first stop the workers:
Tip
Using the --spec option with the merlin stop-workers command will tell Merlin to only stop workers from a specific YAML spec
We stopped these tasks from running but if we were to run the workflow again (with 100 samples instead of 10), we would continue running the 10 samples first! This is because the queues are still filled with the previous attempt's tasks. This can be seen with:
We need to purge these queues first in order to repopulate them with the appropriate tasks. This is where we use the merlin purge command:
Now we are free to repopulate the queues with the 100 samples. In our terminal window that's not designated for our workers, we'll queue up tasks again, this time with 100 samples:
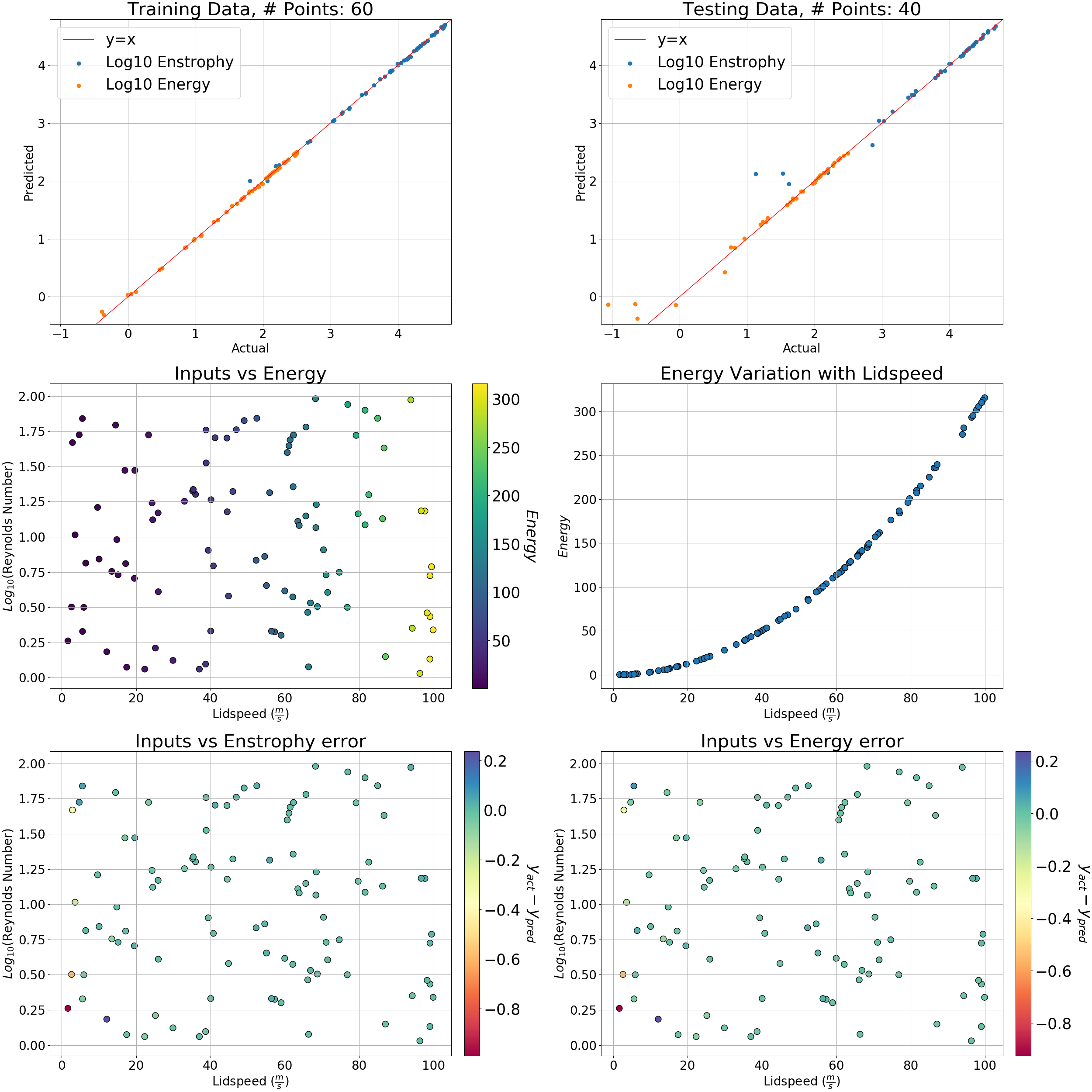
Then in our window for workers, we'll execute:
To see your results, look inside the learn output directory. You should see a png that looks like this: